Invention Summary:
Currently, patients with ER+ breast cancer are administered tamoxifen without considering their potential response to this therapy. Thousands of women take the tamoxifen only to discover that their tumors do not respond or that they develop resistance over time. The ability to screen patients ahead of the therapy administration can improve disease course, management an outcome as patients at risk of resistance can be offered alternative regimens.
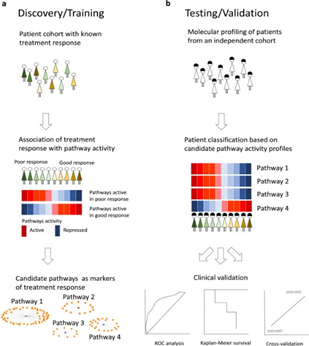
Rutgers Researchers have identified molecular pathways and their read-out genes as markers of tamoxifen resistance in ER+ breast cancer patients. Through the association of pathway activity and response to tamoxifen, five biological pathways were identified as markers of tamoxifen resistance and research demonstrated their ability to predict the risk of tamoxifen resistance in two independent patient cohorts. These pathways were shown not to be markers of aggressiveness and to outperform known markers of tamoxifen response.
For adoption into clinic, a list of pathway read-out genes (i.e., 5 genes) and their associated scoring system were created to assigns a risk of tamoxifen resistance for new incoming patients.
Advantages:
- Improves the therapeutic management of breast cancer patients and builds a foundation for personalized therapeutics for patients with advanced malignancies.
- Superior performance for predicting tamoxifen response when compared to other markers (in literature)
- In the long term, can be employed to improve clinical decision-making, patient survival, and cancer management.
Market Applications: The identified pathways and their read-out genes can be utilized to prioritize patients who would benefit from tamoxifen treatment and patients at risk of tamoxifen resistance that should be offered alternative regimens.
Intellectual Property & Development Status: Patent Pending. Software available. Intellectual property available for licensing and/or research collaboration.